Citation
Orvis J, et al. Nat Methods. 2021 Jun 25.
doi: 10.1038/s41592-021-01200-9
PMID: 34172972
Pancreatic ductal adenocarcinoma (PDAC) arises from heterogeneous precursor lesions, including intraductal papillary mucinous neoplasms (IPMNs), but the features distinguishing indolent from progressive lesions remain unclear. We performed an integrative analysis of transcriptomic, genomic, and microenvironmental profiles of IPMNs to define multi-omic phenotypes. Using transfer learning, we projected IPMN-derived transcriptional programs onto spatial transcriptomic datasets from IPMNs and pancreatic intraepithelial neoplasias (PanINs). We identified two major phenotypes: one associated with cancer-associated fibroblasts and epithelial-to-mesenchymal transition, shared across IPMN, PanIN, and PDAC; and a second, glycolysis-enriched phenotype with a unique somatic variant profile specific to IPMN. Spatial mapping further revealed grade-specific enrichment of transcriptional programs and distinct interactions with stromal and immune subtypes, underscoring the role of the precancer microenvironment in progression. These findings establish multi-omic phenotypes that unify genetic, transcriptional, and microenvironmental heterogeneity, providing a framework for distinguishing progressive from indolent precancers and a web-based public atlas for future exploration of these data and transcriptional phenotypes.
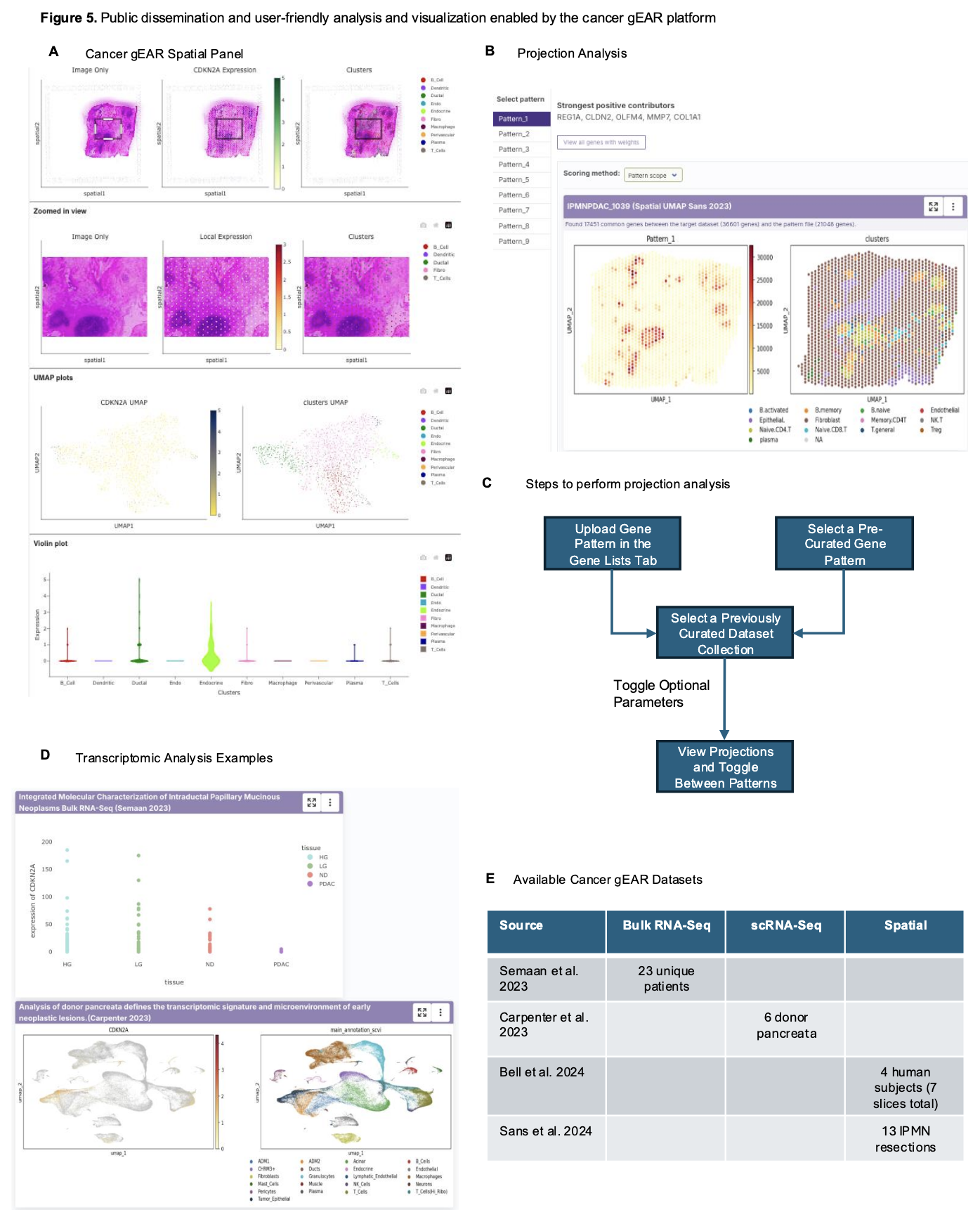
Public dissemination and user-friendly analysis and visualization enabled by the cancer gEAR platform
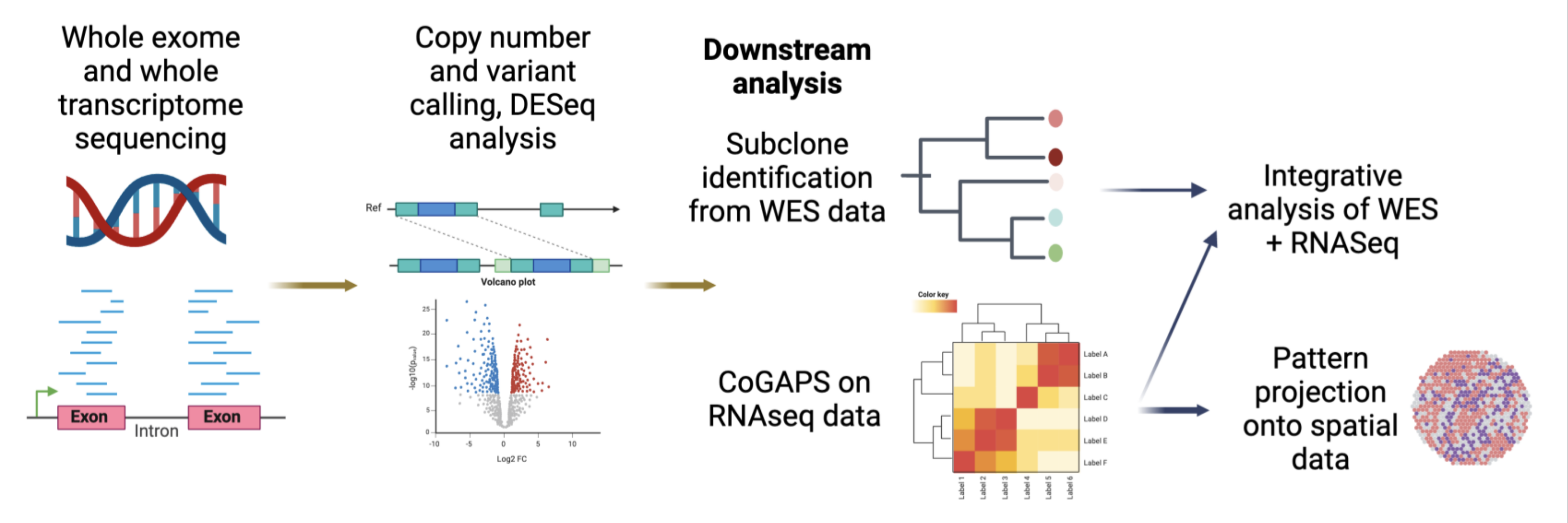
Semaan IPMN paper workflow
Workflow defining transcriptional patterns from bulk RNA-seq data of IPMN lesions from Semaan et al, 2023. CancerGear enables further projection of these signatures across pancreatic precancer datasets, including single-cell and spatial transcriptomics, to evaluate the impact of the microenvironment on these states and the occurrence in different subtypes of lesions. This software also allows for exploration of PanIN signatures defined in Bell et al, 2024, custom gene signatures, and gene expression analyses across collated cohorts of pancreatic precancer data.
IPMN Spatial (Sans 2023): Paper
PanIN Spatial HG and LG (Bell 2024): Paper
Papillary Mucinous Neoplasms (Semaan 2023): Paper
Early Neoplastic Lesions (Carpentar 2023): Paper
CoGAPS_Semaan: Noller et al 2025 - Transcriptional subtypes defined from epithelial cells in IPMN data from Semaan et al, 2023.
CoGAPS_PDAC: Guinn et al 2024 - Transcription subtypes defined for PDAC epithelial cells Guinn et al, 2024 and analysed in PanINs Bell et al, 2024.
Each cell in the table links to the projection of one of the Cancer gEAR data collections described
above into one of the pattern signatures described above, allowing the exploration of each of the pattern
signatures as they change across the samples in the Cancer gEAR data
collections.
The first column of links can be used to view the expression of individual genes across
the data collections, and uses CDKN2A as an example gene to explore its expression patterns.
| Pattern signatures | |||
|---|---|---|---|
| Dataset collections | Individual gene links | CoGAPS_Semaan | CoGAPS_PDAC |
| IPMN Spatial (Sans 2023) | LINK | LINK | LINK |
| PanIN Spatial HG and LG (Bell 2024) | LINK | LINK | LINK |
| Papillary Mucinous Neoplasms (Semaan 2023) | LINK | LINK | LINK |
| Early Neoplastic Lesions (Carpentar 2023) | LINK | LINK | LINK |